Search
Specific Labeling, Enrichment and Quantitation of S-nitrosylated Peptides Using iodoTMT Reagents
Measure protein and peptide cysteine modifications using multiplex quantitative mass spectrometry
by Ryan D. Bomgarden, Ph.D.; Eric L. Hommema, Ph.D.; Rosa I. Viner, Ph.D.1; Zhe Qu, Ph.D.2; Zezong Gu, Ph.D.2; John Rogers, Ph.D. - 08/19/13
S-nitrosylation is a post-translational modification (PTM) that regulates numerous processes, including cellular proliferation, apoptosis, smooth muscle relaxation, neurotransmitter release, and differentiation (Ref.1). During S-nitrosylation, nitric oxide radicals react with free cysteine thiols to produce an S-NO adduct that alters protein activity. Unlike other PTMs such as phosphorylation and acetylation, S-nitrosylation does not appear to have a clear consensus site or motif that can be used to predict which proteins are involved in S-nitrosylation signaling. In addition, due to the highly labile nature of S-NO adduct bonds, S-nitrosylated proteins are difficult to detect and accurately quantify using traditional antibody-based methods.
In 1998, Jaffrey et al. described the biotin switch assay, which used HPDP-biotin (Part No. 21341) to detect S-nitrosylated proteins (Ref.2). In this assay, free sulfhydryls are chemically blocked, whereas nitrosylated cysteines are selectively reduced and biotinylated for streptavidin-based enrichment and/or detection. However, the biotin labeling chemistry used is in this method is reversible, preventing protein S-nitrosylation site identification by mass spectrometry (MS).
Thermo Scientific Iodoacetyl Tandem Mass Tag™ (iodoTMT™) Reagents are a set of isobaric compounds that enable multiplexed quantitation of cysteine-containing proteins using tandem mass spectrometry. The iodoTMT reagents can be used to label up to six different sample conditions which can be combined into a single MS analysis. Unlike pyridyldithiol (HPDP) reagents, iodoTMT Reagents irreversibly label free thiols which allows for labeling under reducing conditions. In this study, we used iodoTMT reagents instead of HPDP-biotin for classic S-nitrosylation switch labeling to identify protein S-nitrosylation sites and quantify changes in S-nitrosylation PTMs by mass spectrometry.
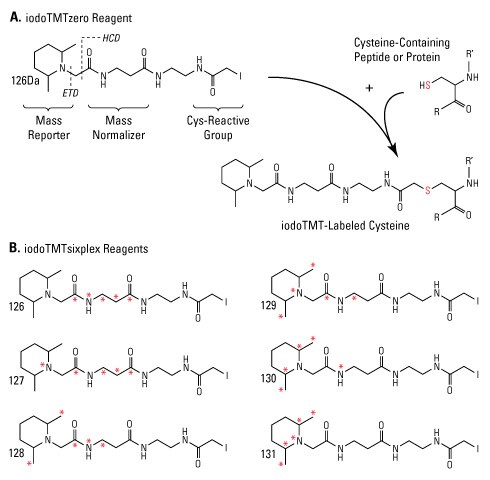
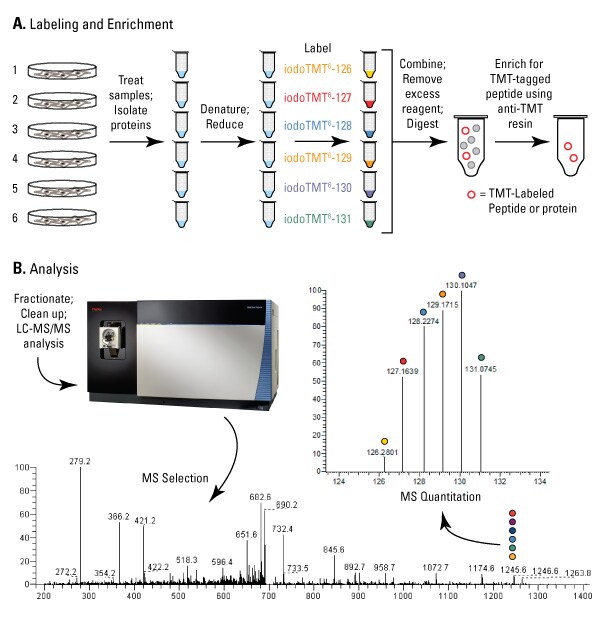
We developed iodoacetyl TMT™ (iodoTMT) reagents to irreversibly label sulfhydryls of cysteine-containing peptides for multiplex quantitation by LC-MS. The iodoTMT reagents are a set of isobaric compounds with the same mass and chemical structure (i.e., isotopomeric) composed of a cysteine-reactive iodoacetyl group, a spacer arm and a mass reporter (Figure 1). The reagent set enables up to six different protein samples prepared from cells or tissues to be labeled in parallel and then combined for enzymatic digestion, enrichment and LC-MS analysis. For each sample, a unique reporter mass (i.e., 126-131Da) in the low-mass region of the MS/MS spectrum is used to measure relative protein expression levels during peptide fragmentation and tandem mass spectrometry (Figure 2, Ref 3).
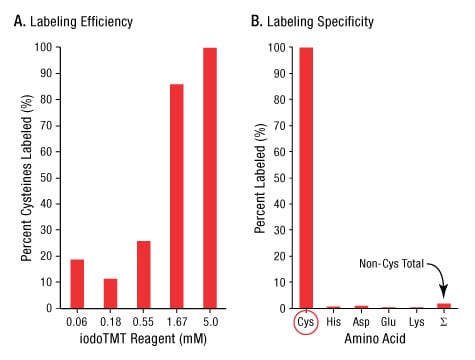
Compared to pyridyldithiol-based reagents such as HPDP-biotin or cysTMT reagents (Ref 4), iodoTMT reagents label proteins and peptides via an irreversible thioester bond. This allows for a simpler sample preparation workflow since samples can be labeled under reducing conditions without the need to remove reducing agent. To determine the optimal concentration of iodoTMT reagents in labeling reactions, we treated bovine serum albumin (BSA) with increasing amounts of tag in the presence of reducing agents. As shown in Figure 3, reactions containing at least 5mM iodoTMT reagent provided complete labeling of all BSA cysteines. Off-target (non-cysteine) labeling was less than 5%, even with an excess (10mM) of iodoTMT reagent. This demonstrates that the reactivity and specificity of iodoTMT reagents is similar to iodoacetamide, which is frequently used to block cysteines for MS analysis.
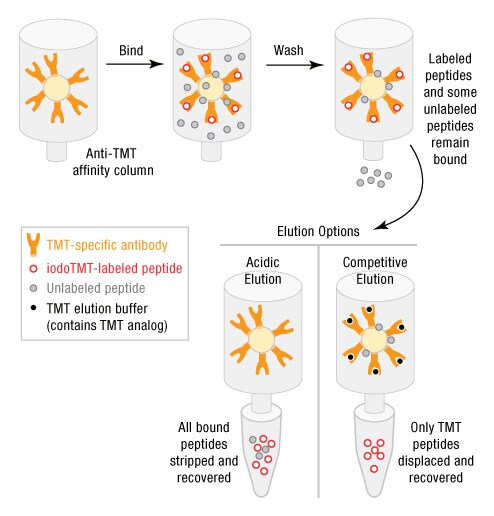
Cysteine is one of the least abundant amino acids, composing only 2.3% of the amino acids of the human proteome. Because of the relatively low abundance of cysteines in tryptic peptide sequences, it is necessary to enrich cysteine-containing peptides before MS analysis in order to identify and quantify labeled peptides (Ref 5). Previously, we developed Immobilized Anti-TMT Resin (Part No. 90076) for capture of TMT-labeled proteins and peptides, as well as an acidic buffer for peptide elution. Although the anti-TMT resin has high specificity for TMT-labeled peptides, the small amounts of unlabeled peptides that non-specifically bind to the resin also elute using the acidic elution buffer. Recently, we developed an improved TMT Elution Buffer (Part No. 90104) for competitive elution of TMT-labeled peptides (Figure 4). An advantage of competitive elution is less contamination of enriched samples with non-specific peptides.
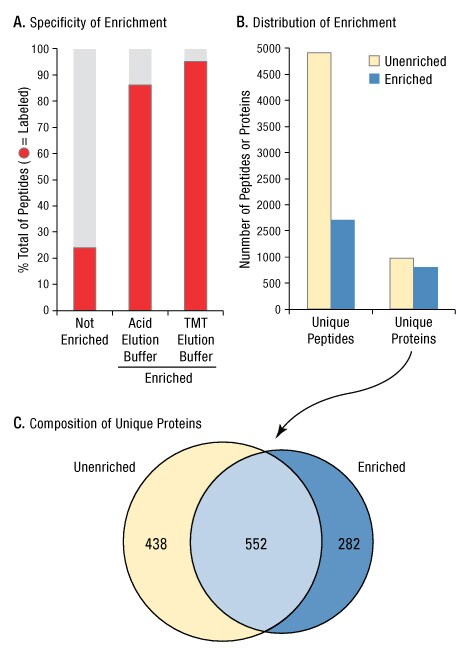
The benefits of enrichment for analysis of cysteine-containing proteins are demonstrated by the data presented in Figure 5. In the absence of iodoTMT labeled-peptide enrichment, only 23% of peptides from an A549 cell lysate digest were modified with an iodoTMT tag. (As demonstrated in Figure 3, it can be assumed that nearly all cysteine-containing peptides are labeled, but only a fraction of total peptides contain cysteines.) In contrast, after anti-TMT peptide enrichment and elution, 95% of identified peptides contained an iodoTMT-modified cysteine (Figure 5A). Although fewer total peptides were identified post enrichment (4916 vs. 1709), the number of proteins identified were similar (973 vs. 814) (Figure 5B). Most importantly, the overlap in composition of proteins identified before and after enrichment using the anti-TMT resin and TMT Elution Buffer was less than 50% (Figure 5C). This demonstrates the low abundance of cysteine peptides, which results in a unique subset of proteins identified by MS analysis upon enrichment.
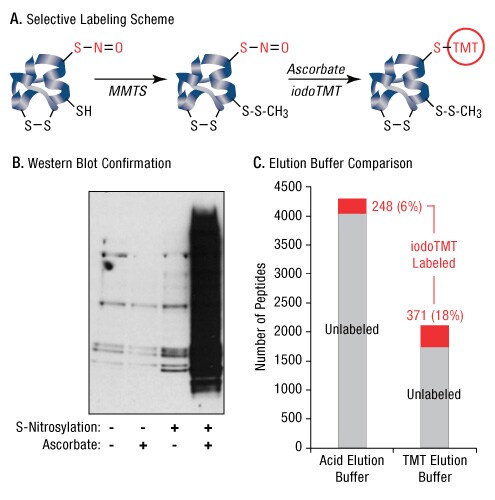
In addition, being one of the lowest abundant amino acids in the proteome, cysteine has unique biochemical properties which allow for a variety of post translational modifications (PTMs) including disulfide bridges, palmitoylation and S-nitrosylation. One method to detect and measure protein S-nitrosylation is a switch assay in which S-nitro groups are replaced with a chemical tag. We used a modified switch assay (Ref 6) to replace S-nitro groups with an iodoTMT tag. In this assay, free sulfhydryls are chemically blocked with MMTS, whereas nitrosylated cysteines are selectively reduced with ascorbate to generate a new free thiol for iodoTMT labeling (Figure 6A). IodoTMT labeling of S-nitrosylated proteins generated using the nitric oxide donor S-nitrosoglutathione was highly specific as shown by Western blot using an anti-TMT antibody (Figure 6B). MS analysis of iodoTMT-switch labeled peptides after anti-TMT enrichment and competitive elution using TMT Elution Buffer resulted in the identification of 371 unique S-nitrosylated peptides. This represents a 50% increase over the number of unique S-nitrosylated peptides recovered by enrichment with acidic elution (248) (Figure 6C).
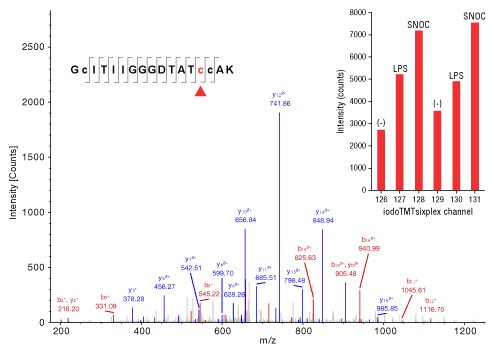
Besides enabling increased identification of low abundant cysteine-containing peptides and site-specific localization of S-nitro PTMs, iodoTMT reagents can also be used to quantify relative changes among six different samples or treatments. To demonstrate this, we performed the following experiment involving 2 x 3 conditions: BV-2 glioma cells were untreated or treated with lipopolysaccharide (LPS) or S-nitrosocysteine (SNOC) for 20 hours to induce S-nitrosylation. Duplicate samples from each condition were selectively labeled with iodoTMT6plex reagents using the modified S-nitrosylation switch assay and combined before digestion, enrichment and MS analysis. Overall, 139 unique peptides were quantified from 174 iodoTMT-labeled peptides for a total of 97 S-nitrosylated protein measured for all six conditions. One protein which showed increased S-nitrosylation upon stimulation with LPS and SNOC was phosphoglycerate kinase 1 (Figure 7). Although the S-nitrosylated peptide identified for this protein contains three cysteines, only one was found to be modified by iodoTMT (red arrow). This exemplifies the power of iodoTMT reagents to help identify, assign and quantify S-nitrosylation sites in peptides with multiple cysteines treated with various S-nitrosylation inducing agents.
Iodoacetyl Tandem Mass Tags (iodoTMT) are novel reagents for specific, irreversible labeling of cysteine residues for multiplexed quantitation of cysteine-containing peptides. Identification and quantification of low-abundance S-nitrosylation modification sites can be accomplished by combining iodoTMT reagent labeling with S-nitrosylation switch chemistry, selective anti-TMT enrichment and competitive TMT peptide elution.
Preparation of iodoTMT-labeled proteins
Proteins and cell lysates were solubilized at 2mg/mL in 50mM HEPES pH 8.0, 0.1% SDS and reduced with 5mM TCEP (Part No. 77720) for 1 hour at 50°C. Reduced proteins were labeled with 5-10mM iodoTMT reagents (Part No. 90100 or 90101) for 1 hr at 37°C protected from light. Excess iodoTMT reagent was removed by acetone precipitation of samples at -20°C for 4-20 hrs. Proteins were digested at 37°C for 4 hrs using trypsin (Part No. 90057) and desalted using C18 spin tips (Part No. 84850) before liquid chromatography (LC)-MS/MS analysis or enrichment.
Selective labeling of S-nitrosylated proteins
Cell lysates were solubilized at 2mg/mL in modified HENS buffer (Part No. 90106). Samples were equally mixed before enrichment of iodoTMT-labeled peptides and LC-MS analysis. Free sulfhydryls were blocked with methyl methane thiosulfate (MMTS, Part No. 23011) for 20 min at room temperature and desalted to remove excess blocking reagent. S-nitrosylated cysteine sulfhydryls were selectively labeled using 0.4mM iodoTMT reagent in the presence of 20mM sodium ascorbate. Excess iodoTMT reagent was removed by acetone precipitation of samples at -20°C for 4-20 hrs. Unlabeled cysteines were reduced and alkylated with 20mM iodoacetamide (Part No. 90034). Proteins were digested at 37°C for 4 hrs using trypsin (Part No.90057) and desalted using C18 spin tips (Part No. 84850) before LC-MS/MS analysis or enrichment.
Enrichment of iodoTMT-labeled peptides
Labeled peptides (25-100μg) were resuspended in TBS at 0.5μg/μL and incubated with 20-100μL immobilized anti-TMT antibody resin (Part No. 90076) overnight with end-over-end shaking at 4°C. After collection of the unbound sample, the resin was washed 4X with 4M Urea/TBS, 4X with TBS, and 4X with water. Peptides were eluted 3X with 50% acetonitrile/0.4% TFA or TMT Elution Buffer (Part No. 90104), frozen, and then dried under vacuum before LC-MS/MS analysis.
LC-MS/MS analysis
A nanoflow high-pressure liquid chromatography system with a Thermo Scientific PepMap C18 column (75 µm ID x 20 cm) was used to separate peptides using a 5%-40% gradient (A: water, 0.1% formic acid; B: acetonitrile, 0.1% formic acid) at 300nL/min over 60 min. A Thermo Scientific LTQ Orbitrap XL ETD linear ion trap mass spectrometer was used to detect peptides using a top-3 CID, -3 HCD experiment for peptide identification and reporter ion quantitation.
Data analysis
MS spectra were searched using Thermo Scientific Proteome Discoverer software v1.4 with SEQUEST™ and Mascot™ programs against a mammalian Swiss-Prot™ database. Modifications included carbamidomethyl, iodoTMTzero (324.42 Da) and iodoTMTsixplex reagent (329.38 Da) for cysteines and oxidation for methionine.
- Hess, D.T. and Stamler, J.S. (2012). Regulation by S-nitrosylation of protein post-translational modification. J. Biol. Chem. 287(7):4411-8.
- Jaffrey, S.R. and Snyder, S.H. (2001). The biotin switch method for the detection of S-nitrosylated proteins. Sci STKE (86)1-9.
- Thompson, A., et al. (2003). Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal. Chem. 75:1895-904.
- Murray, C.I, et al. (2012). Identification and quantification of S-nitrosylation by cysteine reactive tandem mass tag switch assay. Mol. Cell. Proteomics11(2): M111.013441.
- Gygi, S.P., et al. (1999). Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotech.17:994-9.
- Forrester, M.T., et al. (2009). Detection of protein S-nitrosylation with the biotin technique. Free Radic. Biol. Med. 46(2):119-26.
The data in this article were previously presented at the 2012 American Society for Mass Spectrometry annual meeting in a poster titled: Iodoacetyl tandem mass tags for cysteine peptide modification, enrichment and quantitation.
仅供科研使用,不可用于诊断目的。